Coordinated by PAHO and comprised of laboratories from 24 countries in the Region, the COVID-19 Genomic Surveillance Regional Network (COVIGEN) monitors the virus that causes COVID-19 to detect any change in its sequence that may influence its ability to spread, disease severity, vaccine effectiveness, treatments, diagnostics, or other public health and social measures. How does the network work? What is the current distribution of the variants of concern? And why is this network important to help end the pandemic and prepare for the future?
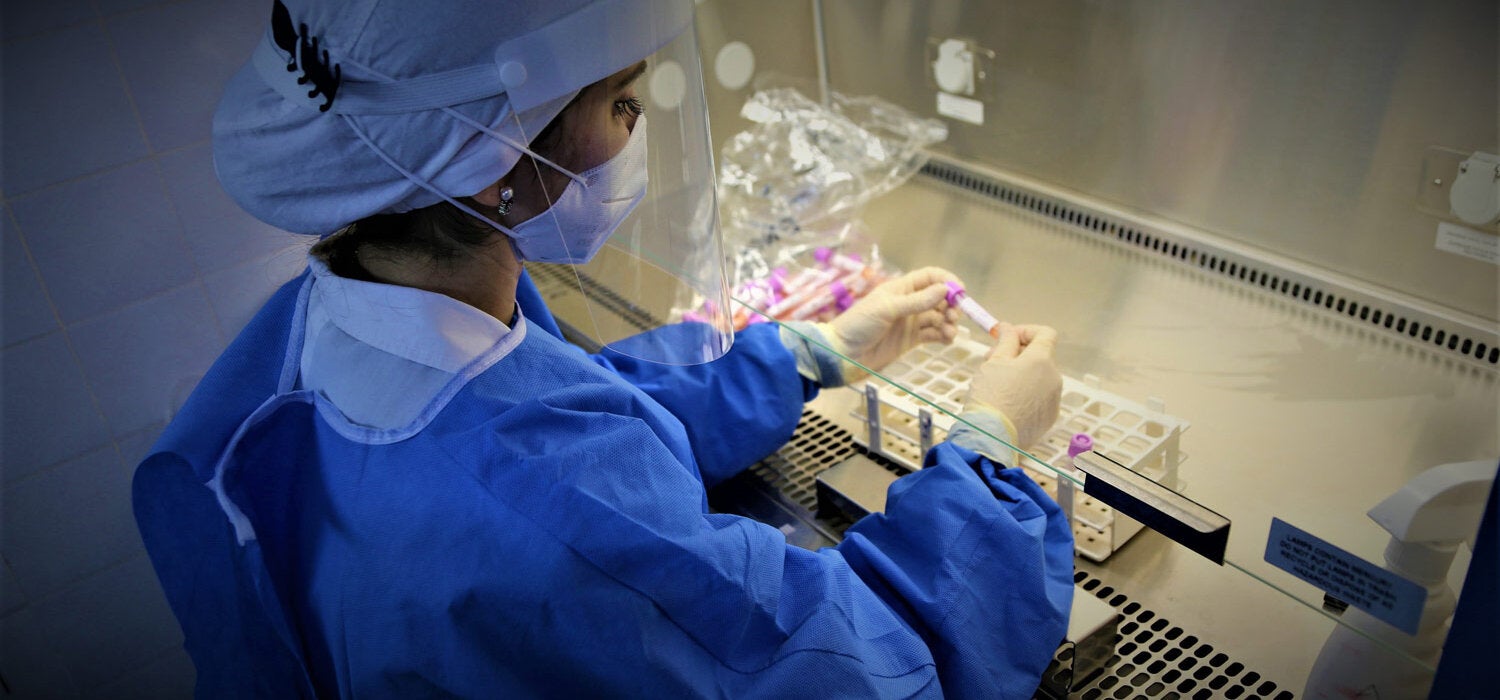
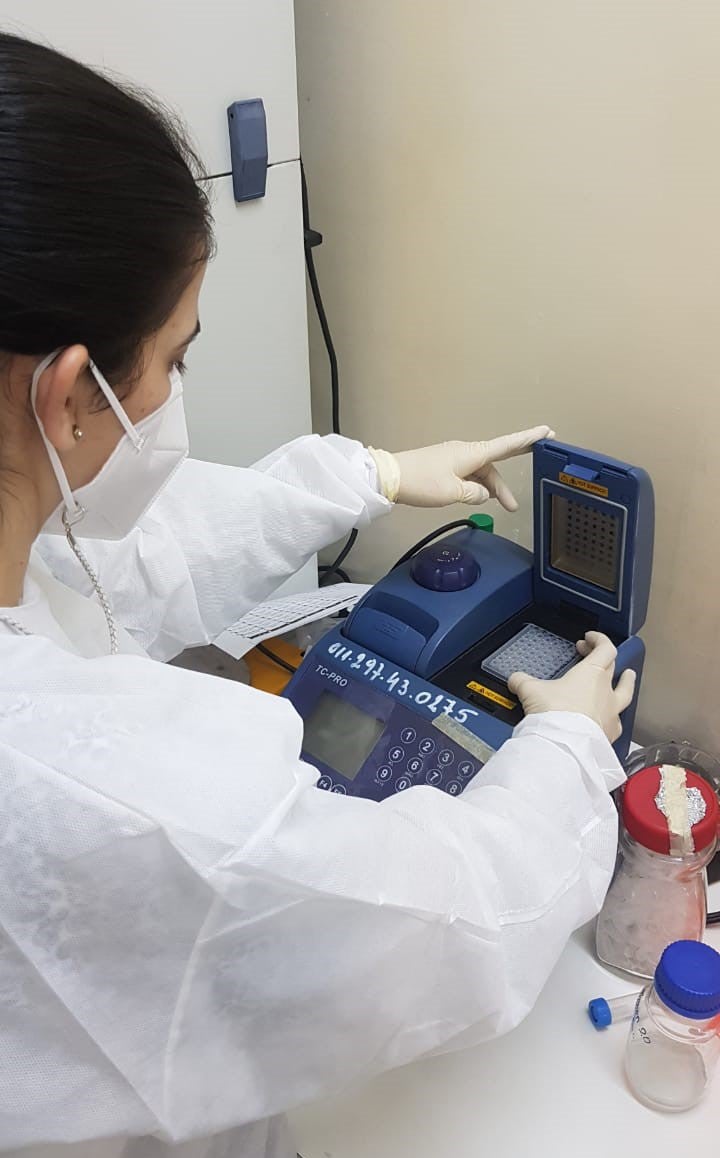
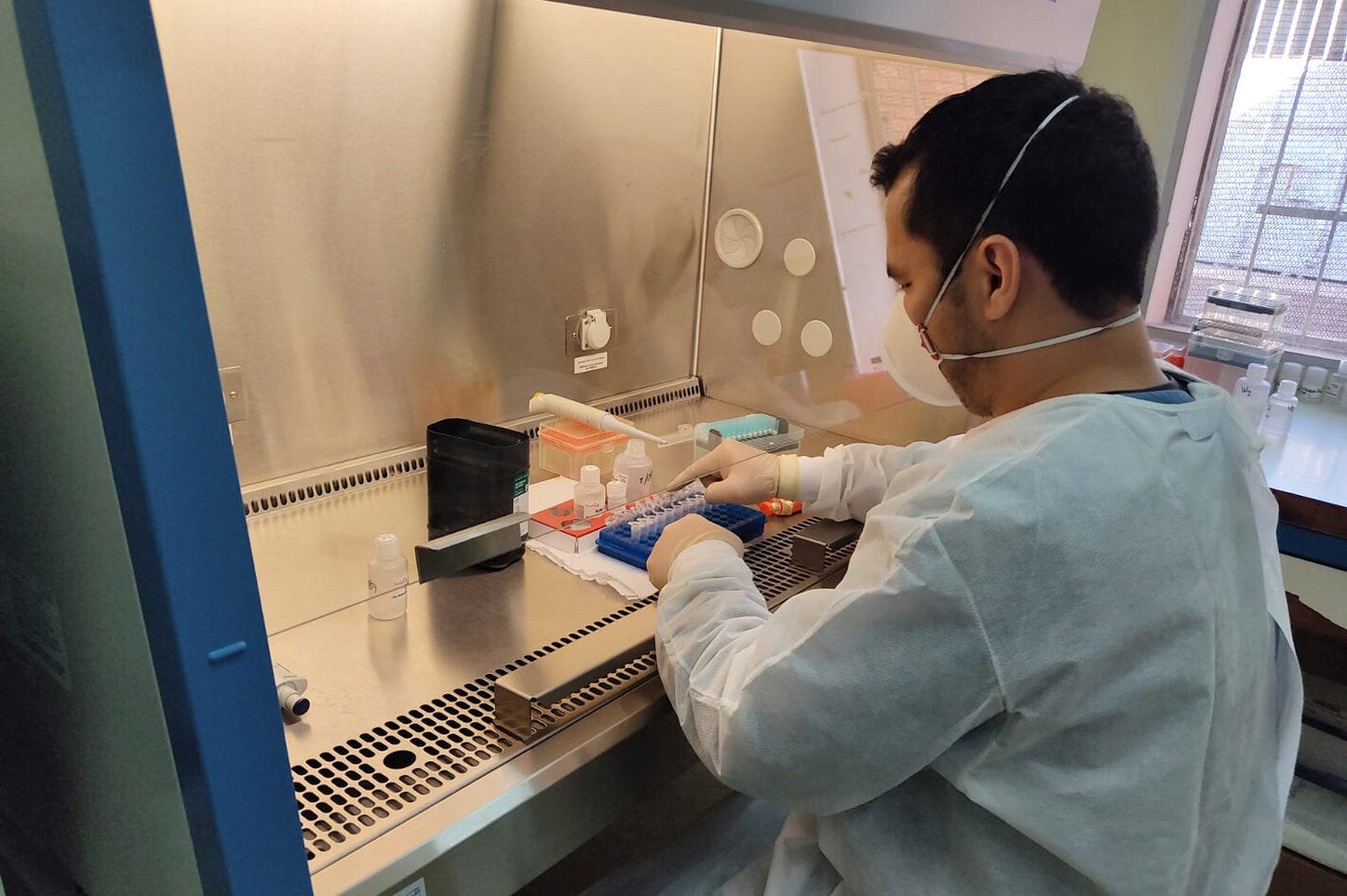
Washington, DC, July 2021 (PAHO) - Fifty samples from people who tested positive for COVID-19 depart from Santo Domingo airport in the Dominican Republic for Rio de Janeiro. Their destination is the Oswaldo Cruz Foundation (FIOCRUZ) laboratory for genomic sequencing of the SARS-CoV-2 virus. The results of these strategically selected samples representing different regions of the country will produce an overview of circulating variants and provide guidance for public health measures to fight the pandemic. Lack of established capacity for genomic sequencing and monitoring of the virus that causes COVID-19 has not a been an issue for the countries of the Americas since the Pan American Health Organization (PAHO) created the COVID-19 Genomic Surveillance Regional Network (COVIGEN) in March 2020, comprised of public health laboratories from 24 countries in the Region.
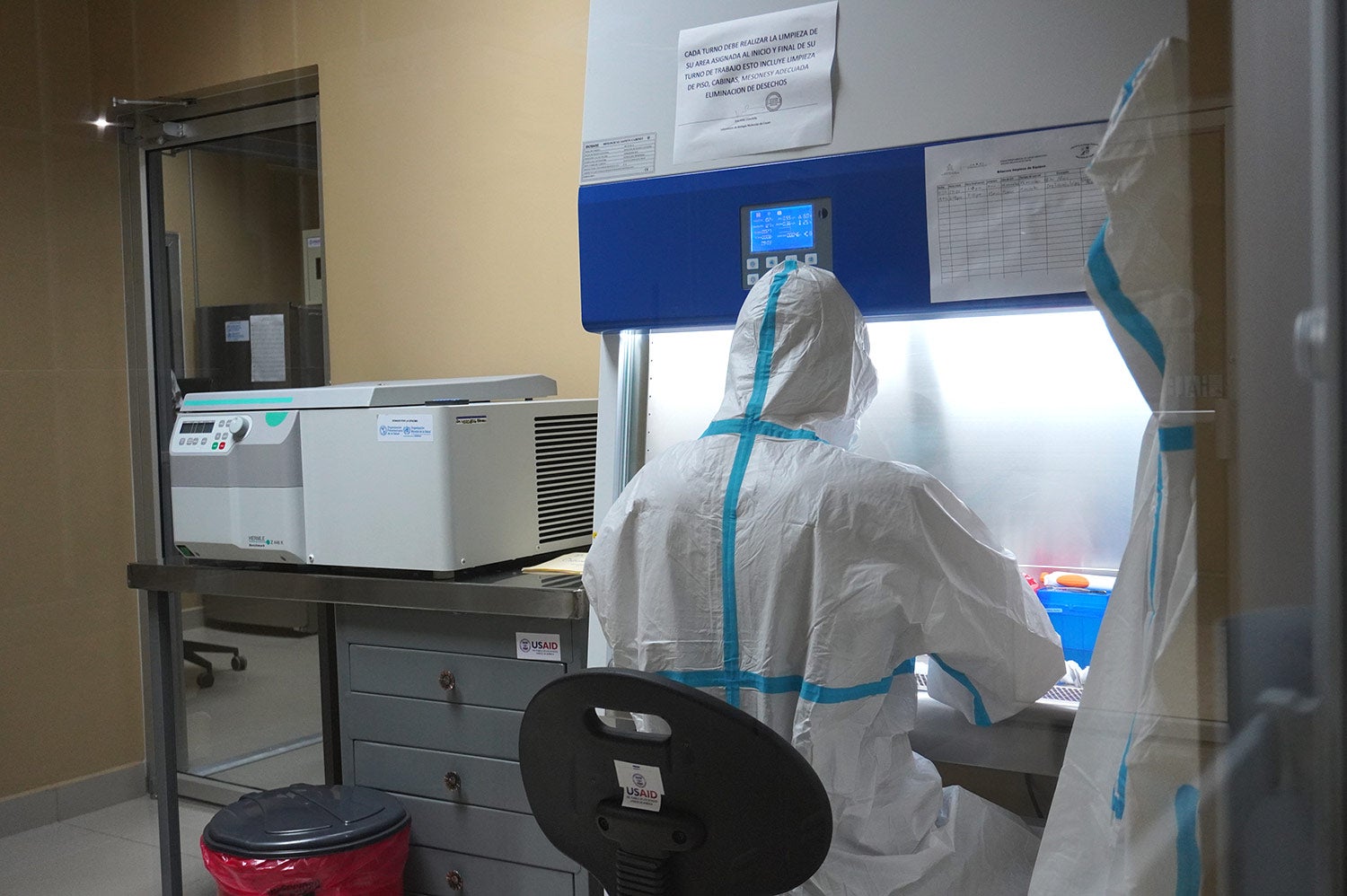
Genomic sequencing of viruses is a relatively new and complex process requiring large investments in high-cost equipment, trained human resources, reagents, and other supplies. Not every country in the world and the region has that capacity in their public health laboratories," said Jairo Méndez Rico, PAHO advisor on emerging viral diseases.
Thirteen countries participating in the network have limited capacity or are in the process of expanding their sequencing capabilities. Those countries are sending their samples to one of the six regional reference laboratories, located in Brazil, Chile, Mexico, Panama, Trinidad and Tobago, and the United States. These efforts made it possible to detect variants of concern for the first time in Bolivia, the Dominican Republic, Guatemala, Haiti, and Honduras.
Using PAHO resources and an exchange between participating laboratories, the network also strengthened capacity in ten countries that had some prior sequencing capability. PAHO supports the project by standardizing screening protocols for variants of concern, collaborating in results analysis, donating reagents, and training laboratory staff.
SARS-CoV-2 surveillance is key for detecting any change in the virus' sequence that may influence its spread, COVID-19 severity or vaccine effectiveness, treatments, diagnosis, or other public health and social measures. "Tracking the virus is vital to inform countries and the public in a timely manner about possible changes that may complicate our response to the virus and transmission prevention," said Méndez Rico.
The network's efforts not only increased the amount, but also the quality of sequencing data at the regional level while contributing to global knowledge about the virus. Although data was initially reported only from Canada and the United States, the Region currently reports data from 49 countries. "This gives us an idea of what is happening across the Region and how these viruses are spreading," the PAHO advisor said.
"There is no exact number of samples per country that must be sequenced to monitor SARS-CoV-2, but it's not about sequencing simply for the sake of sequencing," Méndez Rico explained. "Surveillance is effective when the samples being analyzed represent what is happening in a country and are associated with the epidemiology of the disease, travelers, or an outbreak or cluster," he added.
Using the Global Initiative on Sharing Influenza Data (GISAID) to immediately share sequences of the COVID-19 virus worldwide is also key to public health actions, improving diagnostic protocols, and generating information for vaccine development. "Both public and private laboratories must contribute to knowledge of COVID-19 and response to the disease," he stressed.
According to the PAHO's advisor on emerging viral diseases, the Genomic Surveillance Regional Network has planted the seed for "routine genomic surveillance in the Americas when the pandemic ends," not only for other circulating viruses, such as dengue, arenaviruses, and influenza, but also new viruses or pathogens, allowing us to be better prepared for the next pandemic.
Variants of interest and concern in the Americas
Mutation is a natural part of the evolution of viruses. Since its initial genomic characterization, SARS-CoV-2 has mutated and evolved. A mutated virus is considered a variant of the original virus, and although most mutations have no impact, some may be advantageous to the virus and result in higher transmissibility or greater ability to evade both natural and vaccine-generated immune response.
WHO has designated a group of variants of interest and concern, updated based on the emergence and identification of new variants that have a significant and demonstrated impact on global public health. Country authorities may also designate other variants of local interest or concern.
A variant is considered of interest (VOI) if it is different than the original virus and has been identified as a cause of community transmission/multiple cases/clusters of COVID-19 or has been detected in several countries. As of July 2021, WHO has identified four global variants of interest (eta, iota, kappa, and lambda), two of which have been detected in the Americas (lambda and iota).
A VOI becomes a variant of concern (VOC) if it is associated with increased transmissibility or harmful changes in COVID-19 epidemiology such as increased virulence or change in clinical presentation, or decreased effectiveness of control measures, diagnostic tools, vaccines, and treatments. As of July 2021, WHO has identified four global variants of concern: alpha, beta, gamma, and delta.
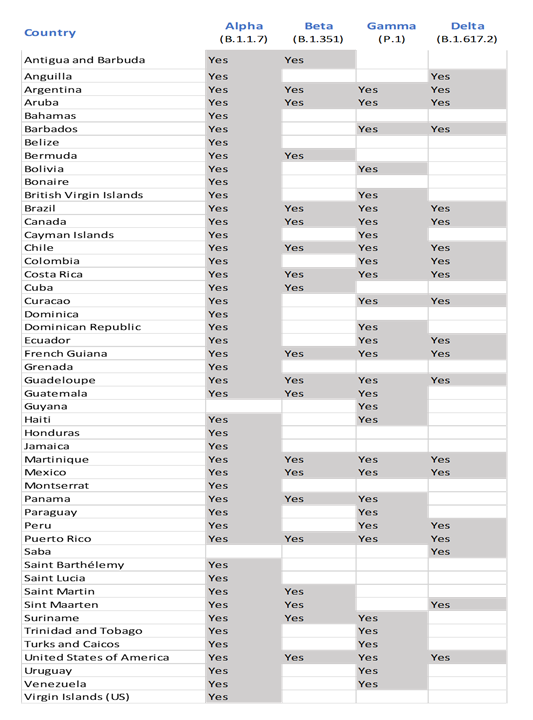
The Genomic Surveillance Regional Network has identified variants of concern in 49 countries of the Americas. Alpha was detected in 45 countries, beta in 20, gamma in 31, and delta in 20. All four VOCs have been identified in 12 countries in the Region.
So far, the available evidence has shown that some variants of concern have greater ability to transmit and replicate; however, there is insufficient evidence to confirm increased aggressiveness or severity, and their impact on reinfections or ability to evade the immune response has yet to be determined," said Méndez Rico. "The information shows that existing vaccines protect against all variants currently circulating globally," he added.
In practical terms, the PAHO expert said, it's still the same virus and the measures to prevent transmission haven’t changed: vaccination, physical distancing, wearing masks, avoiding crowds, and infection control through testing, contact tracing, isolation, and quarantine.
Méndez Rico explained that viruses are organisms that evolve. He said viruses need people's cells to multiply. "When evolving it seeks higher transmissibility to infect more people; however, the virus does not want to kill the person, because then, where would it multiply?" The specialist informed that, in general, viruses, including SARS-CoV-2, evolve to become more transmissible, but not necessarily more aggressive, lethal, or severe.
Variants of Concern in the Americas as of 28 July 2021
Ministry of Health of Paraguay performs SARS-CoV-2 virus sequencing for the first time
Becoming familiar with the genetic sequence of SARS-CoV-2 to monitor or detect its variants is no easy task, but the Ministry of Health of Paraguay is now up to the task.
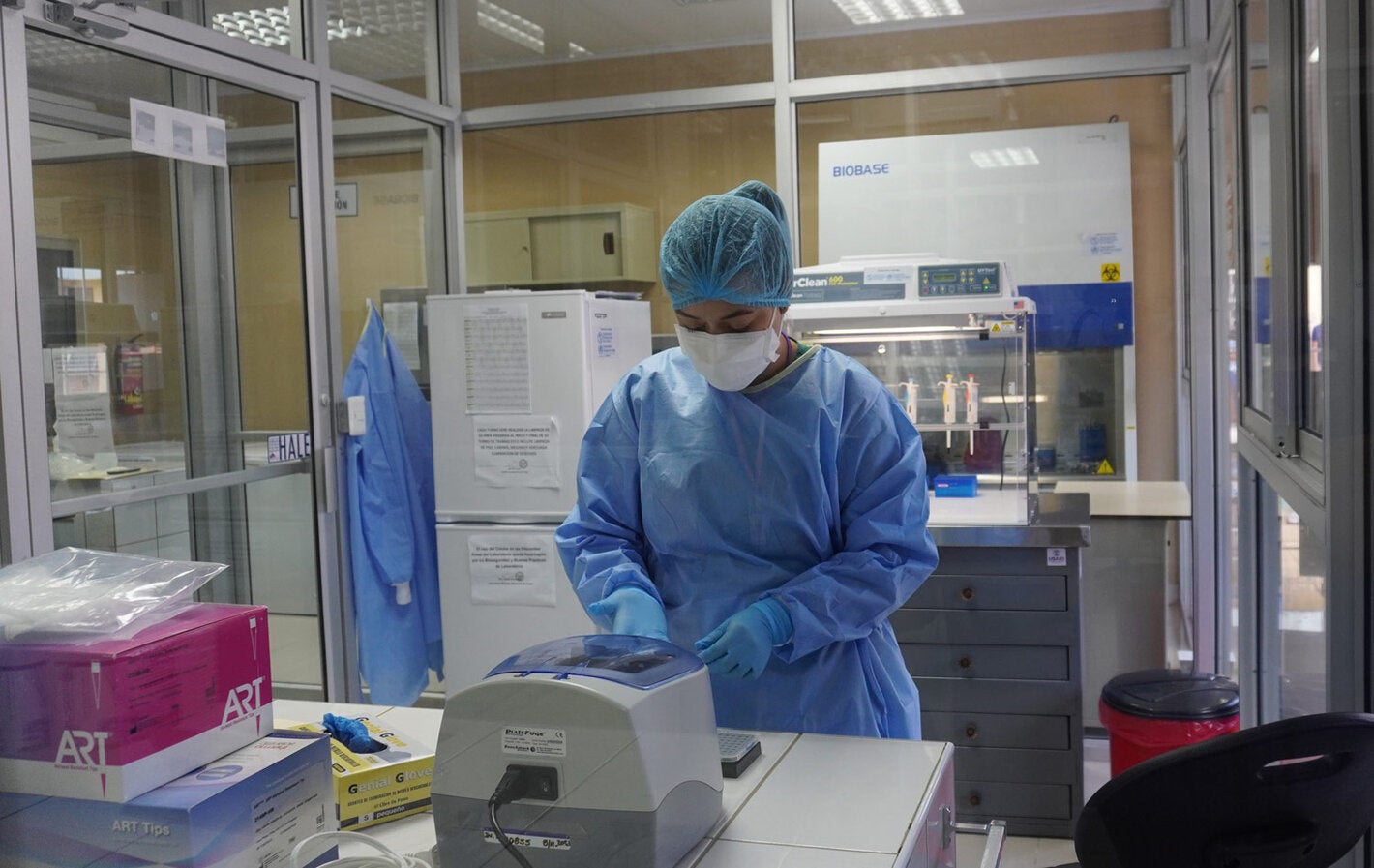
PAHO and the Genomic Surveillance Regional Network have supported the country by strengthening equipment capacity at the Central Public Health Laboratory of Paraguay, which has begun to generate its own sequences of the SARS-CoV-2 virus.
The Ministry had one of the two machines capable of sequencing in Paraguay, but that was not enough. It needed to build the capacity of technicians to carry out the process and to access reagents. Thus, last April PAHO sent a specialist from the laboratory in Paraguay to Fiocruz, one of the network's reference laboratories located in Rio de Janeiro, where she was trained for a week.
Biochemist Sandra González learned about how to assemble a genomic library, the sequencing process, and bioinformatics analysis to extract lineage and variant information. Upon her return, together with the rest of the laboratory team, they ran several tests, and, after the arrival of all the necessary materials, successfully carried out their first sequencing at the end of July.
PAHO provided supplies and reagents for the tests, which were difficult to obtain due to the high global demand and flight restrictions. The protocol developed and validated by Fiocruz for sequencing was also shared for implementation in the country, and the Gorgas Institute of Panama, another member of the network, supported the country in validating the bioinformatics analysis of the results.
"Doing sequencing is a very important step that will allow us to optimize time to obtain fundamental information in a timely manner," says Cynthia Vázquez, technical director of Paraguay's Central Public Health Laboratory. "The support of PAHO and colleagues in the Network was extremely important," she adds.
Cynthia explains that they will continue to use real-time PCR tests to screen for variants, something that can identify with some probability some of the variants of concern, and that they will focus sequencing on identifying the delta variant of concern and the lambda variant of interest. In addition, she says, "we are going to start training more people, to transfer capacity.". An achievement for Paraguay and for global efforts against the pandemic.
Haiti strengthens variant detection
When Haiti began to record an unprecedented increase in COVID-19 cases in May 2021, it became a priority to know which variants were circulating.
"When we first detected COVID-19, we didn't know exactly which variant it was, but now we know it's alpha and gamma," says Ito Journel, head of molecular biology at Haiti's National Public Health Laboratory, referring to the variants of concern detected in the country through mid-July 2021.
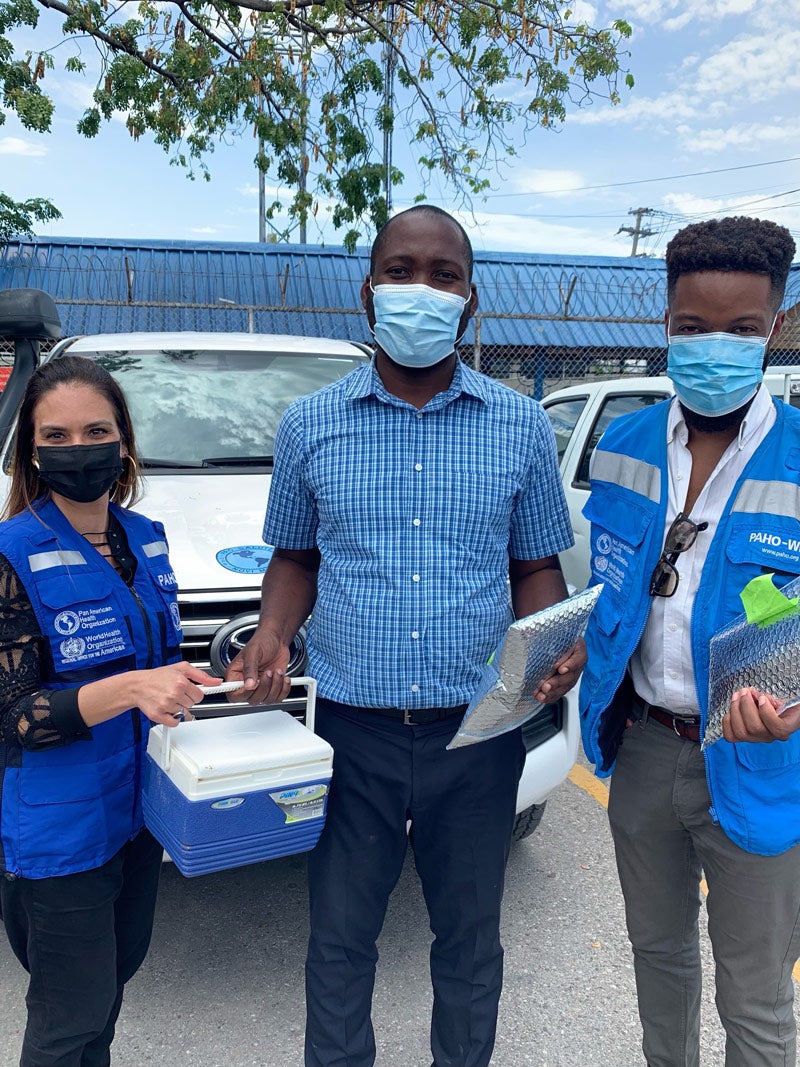
Ito says that PAHO and the COVID-19 Genomic Surveillance Regional Network (COVIGEN) made it possible for samples to reach the Fiocruz laboratory in Rio de Janeiro, Brazil, one of six regional COVIGEN reference laboratories that perform sequencing for countries in the network that lack capacity at the local level. There, the samples were sequenced for the first time in early June and two variants of concern were confirmed.
The capacity for PCR testing, a necessary step prior to sequencing, was also strengthened in Haiti through cooperation with PAHO.
We received technical support, training, supplies, reagents, and financial support from PAHO," Ito explains, adding that even before COVID-19, "PAHO has supported us with two laboratory technicians since 2015 for surveillance of other viruses. When the pandemic started, they provided us with two additional technicians in my lab to do testing more quickly."
With supplies now available, the country can perform initial monitoring and screening of three variants of concern. By participating in the COVIGEN network, Haiti can confirm or detect variants of concern, including delta, by accessing reference laboratories to sequence samples previously selected according to established criteria.
"Now we want to focus on knowing if the delta variant is in the country. We know that there are some cases in the Dominican Republic and Florida. People traveling to Haiti may have brought it here, but we have to send samples to sequence for confirmation," he says, talking about a new process they will do every month.
The Haitian lab technician has been awarded a scholarship for a master's degree at Fiocruz, allowing him to implement sequencing technology in Haiti after completing a two-year program.
He believes that sequencing capacity is very important, especially in a pandemic like this, when reference laboratories may be overburdened with tests for other countries. Establishing capacity in the country also represents an opportunity to expand genomic surveillance for other viruses. "It’s important to have the capacity for genomic surveillance in the country, not just for SARS-CoV-2, but also other viruses that we need to sequence," he says.
While Ito starts his master's degree in Brazil in late August 2021, a team will continue his work in Haiti. But Ito assures that he will always remain in contact with his laboratory by email or text and continue to be involved with SARS-CoV-2 surveillance in his country.
Honduras: Genomic Surveillance Network Helps Confirm Alpha Variant
In June 2021, the National Virology Laboratory of Honduras analyzed samples from COVID-19 positive patients and detected variants of concern through PCR testing. However, lacking the capacity to sequence and confirm the identity of these samples or find other variants that typically evade PCR screening, Dr. Mitzi Castro turned to the Gorgas Institute of Panama, one of the six Genomic Surveillance Regional Network (COVIGEN) reference laboratories. With support from PAHO, the samples arrived in Panama and days later, Gorgas confirmed the presence of the alpha variant of concern in Honduras.
With PAHO support, the Honduran laboratory can perform PCR testing to detect and monitor the alpha, beta, and gamma variants. "We now have the protocol for variants of concern (VOCs) that allows us to screen for three of the four VOC sharing the same mutations, and it works very well," she says.
However, screening is no substitute for sequencing. The protocols and PCR tests for detecting the alpha, beta, and gamma variants do not work for the delta variant, as it does not have the same mutation. As a result, following PAHO's recommendations, "the country sends samples that test negative for the three variants to the reference laboratories in order to identify delta or any other variant by sequencing."
Mitzi Castro highlights PAHO's support in developing criteria to select samples for sequencing, sending samples to COVIGEN network laboratories, delivering reagents to perform PCR tests locally, and providing training on sharing results globally through the GISAID platform.
The lab director says that the analyses performed there have revealed an important issue: "We are seeing that young people, aged 15-48 years old, are more affected by the variants. We are sharing clear and accurate information in the media, so that people are aware and take action to change their behavior and protect themselves."
Other common variants are also circulating in Honduras, but so far, the delta variant has not been confirmed.
Castro assures that if other variants are identified in the country, they will continue "with weekly surveillance, establishing protocols for screening variants, collecting samples and analyzing the data, and sending samples to the reference laboratories for sequencing to detect any other variant of concern." Finally, the specialist emphasizes that: "the lab is the backbone of diagnosis and timely results mean timely actions."
What is genomic sequencing?
The SARS-CoV-2 virus, like other viruses including influenza, is constantly changing. After periods of high circulation, viruses undergo changes in their genomes, producing "mutations" or "variants.".
Genetic sequencing is a laboratory technique that allows the genome of a virus to be read. As each pathogen has a unique genomic sequence, this method can be used to identify novel pathogens (as in the case of SARS-CoV-2).
Studying the evolution of SARS-CoV-2 allows scientists to detect mutations associated with changes in the transmissibility and/or pathogenicity of the virus, or that could reduce the utility of medical countermeasures (diagnostics, vaccines, and therapeutics).
Additionally, following virus mutations over time and space can help to track the spread of the pathogen and support an enhanced understanding of potential transmission routes and dynamics.
Genomic sequencing is used to monitor changes in the virus' genome and collect information about those mutations to determine if the virus has become more transmissible, if existing vaccines are still effective, or if diagnostic tests are still able to detect the changed virus, among other conclusions.
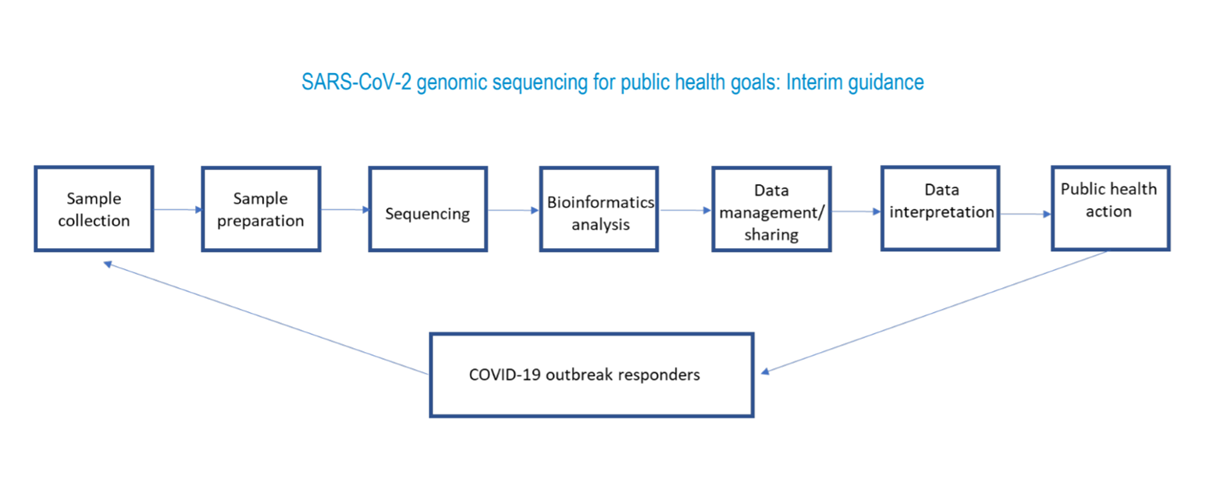
From sampling to public health measures
Collecting samples is the first step in identifying variants circulating in a country. For example, a sample may be taken from a symptomatic person who underwent a PCR test at a health center and tested positive for COVID-19.
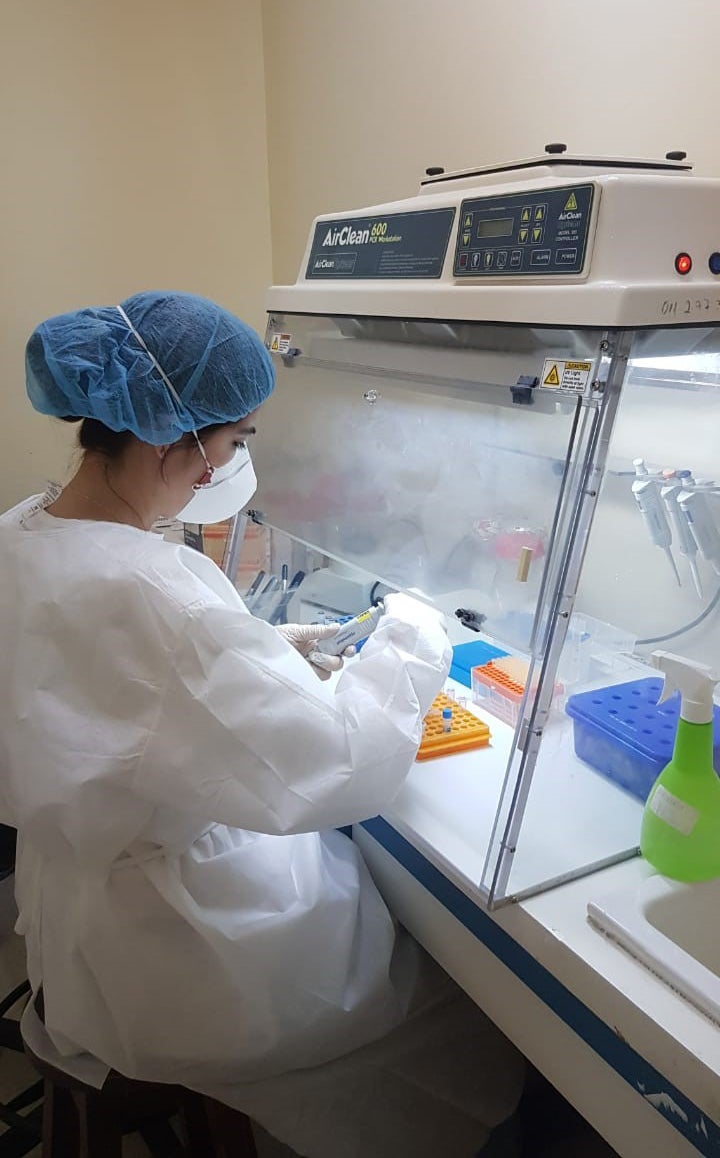
That sample, along with others from the country are sent to a national virology laboratory where they are prepared for sequencing. This involves selecting the material to be sequenced.
Selection should be based on defined criteria not only to ensure quality, but also adequate representation of the variants circulating in the country. Prior to sequencing, the national public health laboratory can perform a series of trials that work like diagnostics, designed to recognize some of the virus' mutations and screen between samples to determine which ones may be alpha, beta, or gamma. Some tests may even detect specific changes in each variant.
However, these trials are considered preliminary and should be supplemented with sequencing.
Although some SARS-CoV-2 viruses in circulation may have specific mutations, screening trials alone are not enough to classify them as variants of concern or interest. For this reason, complete genomic sequencing is required to confirm variants of concern," explains Juliana Leite, a PAHO laboratory and viral disease specialist.
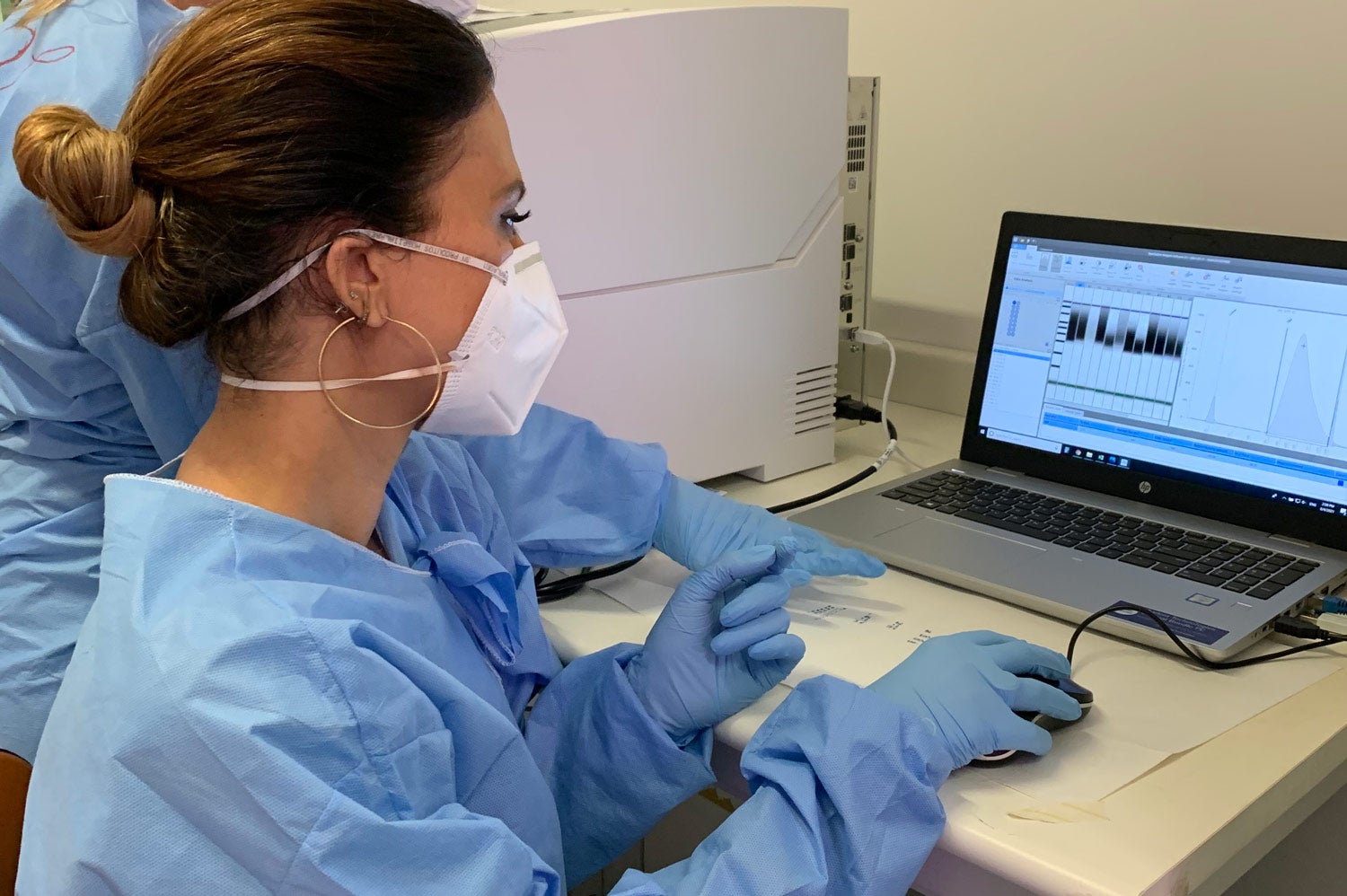
After sequencing, laboratory personnel proceed to bioinformatics analysis. This confirms the presence of alpha, beta, or gamma, and identifies delta or other variants that may arise. This analysis is part of the genomic surveillance process and is performed in countries with adequate capacity or by one of the Genomic Surveillance Regional Network reference laboratories.
After analysis, the data is published in GISAID and is available worldwide. The widespread availability of SARS-CoV-2 sequences, diagnostic procedures, and sequencing protocols and samples has been beneficial to achieving worldwide genomic surveillance of the virus.
Through genomic surveillance of SARS-CoV-2, experts interpret the data to study circulating lineages and variants, including any mutations, and share their analysis with pandemic response authorities to guide public health measures and provide recommendations to the population.